Research Overview
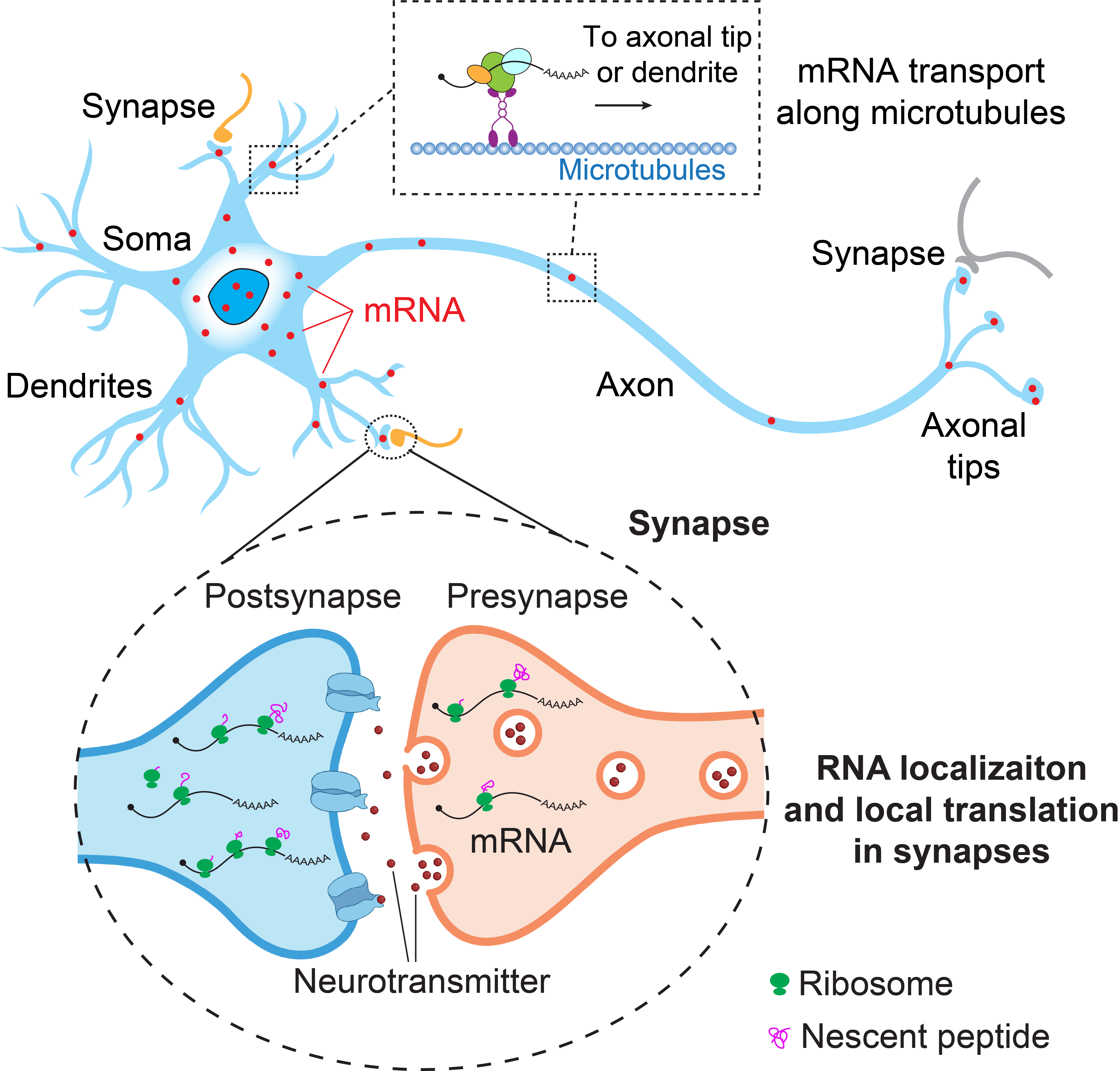
Spatial RNA localization is a key epigenetic mechanism regulating the spatiotemporal gene expression. This mechanism is particularly important in neurons and plays a crucial role in neuronal functions including neurite regeneration and synaptic plasticity which is the cellular basis underlying learning and memory and neurite circuit formation. Aberrant RNA localization has been observed in neurological disorders including Amyotrophic Lateral Sclerosis (ALS), Frontotemporal Dementia (FTD), and Spinal Muscular Atrophy (SMA), and has also been implicated in substance use disorders. However, the regulatory mechanisms, functional roles, and pathological relevance of spatial RNA localization in neurons remain poorly understood, largely due to limited tools. The Han lab aims to address these knowledge caps by applying and developing cutting-edge synthetic biology, chemical biology, and CRISPR technologies to profile, image, and manipulate the spatial transcriptome. Our long-term goal is to develop novel therapeutic strategies to treat neurological disorders including neurodegenerative diseases and substance use disorders via spatial transcriptome engineering.
Research Directions
Direction 1: Engineering RNA-targeting CRISPR systems for studying RNA biology
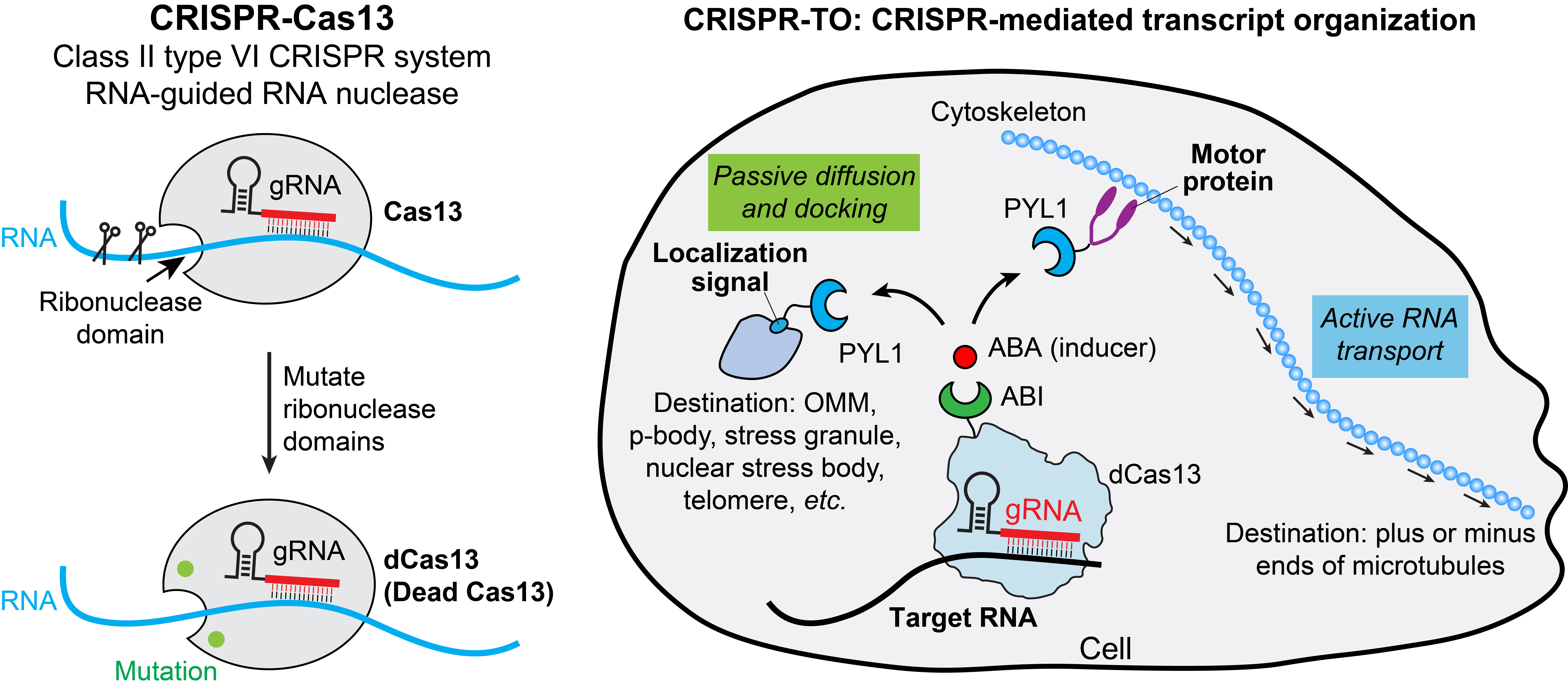
Our lab uses Class 2 type VI CRISPR-Cas13 systems to perturb spatial RNA localization for functional studies. CRISPR-Cas13 is an RNA-guided, RNA-targeting system which uses guide RNA (gRNA) to direct the ribonuclease Cas13 to cleave the target RNA. Mutating the ribonuclease domain of Cas13 creates a catalytically inactive (dead) Cas13 variant (dCas13), which allows targeted RNA binding without RNA cleavage. However, dCas13 is relatively large for in vivo applications and exhibits moderate RNA-binding efficiency. We are actively doing protein engineering to develop better RNA-targeting CRISPR systems and tools for RNA imaging, perturbation, and screening to facilitate the study of RNA biology.
Direction 2: Deciphering the role of RNA localization and local translation in learning and memory
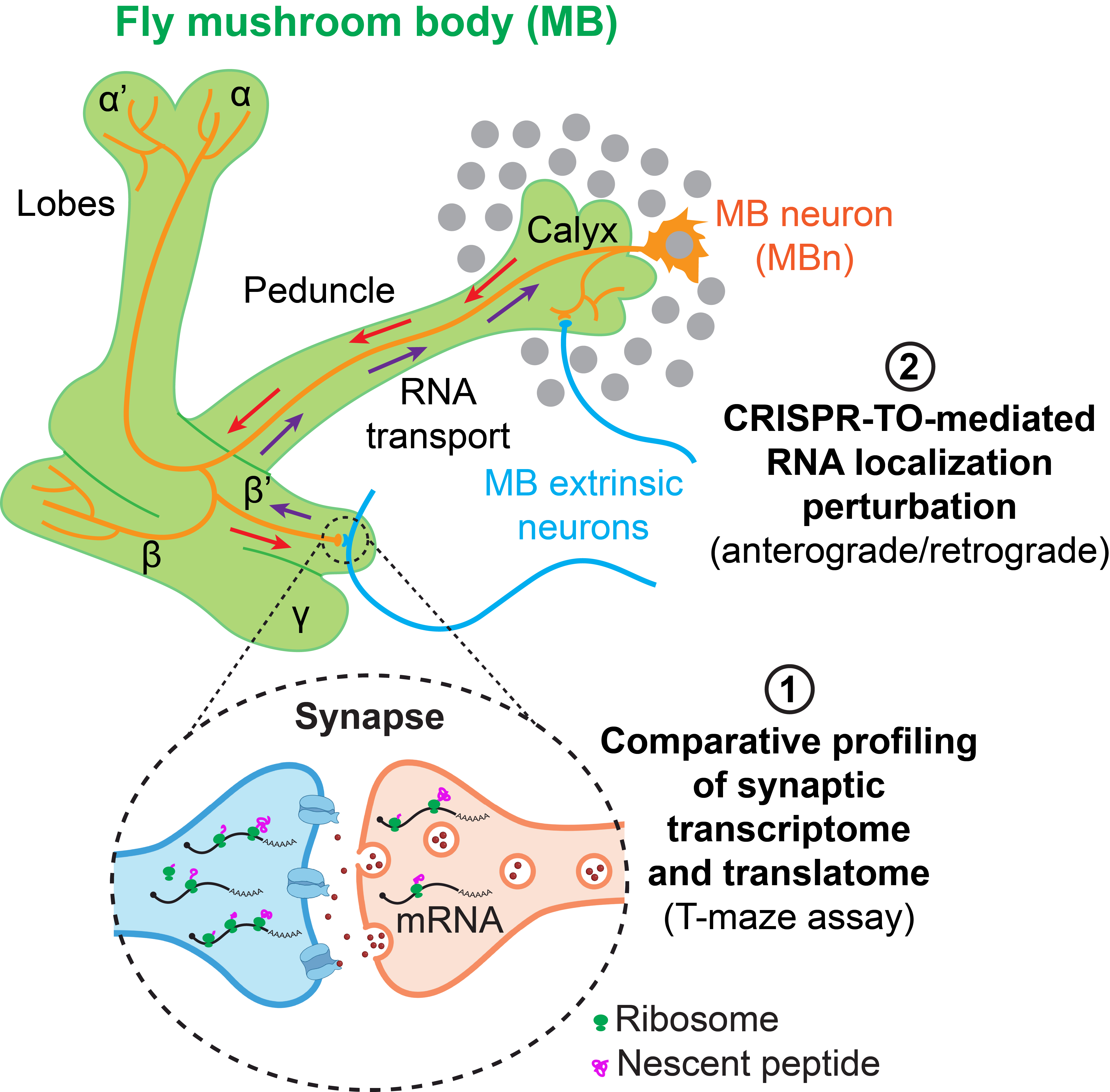
Our lab is interested in understanding how RNA localization and local translation participate in regulating the behavior of learning and memory, including its regulatory mechanisms and functional roles. We use Drosophila as an animal model to study this biological question because it serves as a powerful model for investigating human memory, with highly conserved molecular mechanisms, robust and quantitative behavior assays for learning and memory, and straightforward genetic tractability. We are developing a cutting-edge, multidisciplinary framework integrating comparative synaptic transcriptome and translatome profiling, CRISPR-TO-mediated RNA localization perturbation in Drosophila mushroom body neurons (MBns), and the T-maze olfactory conditioning assay to study this biological question.
Direction 3: Interrogating synaptic RNA localization and local translation in substance use disorders for developing novel therapeutics
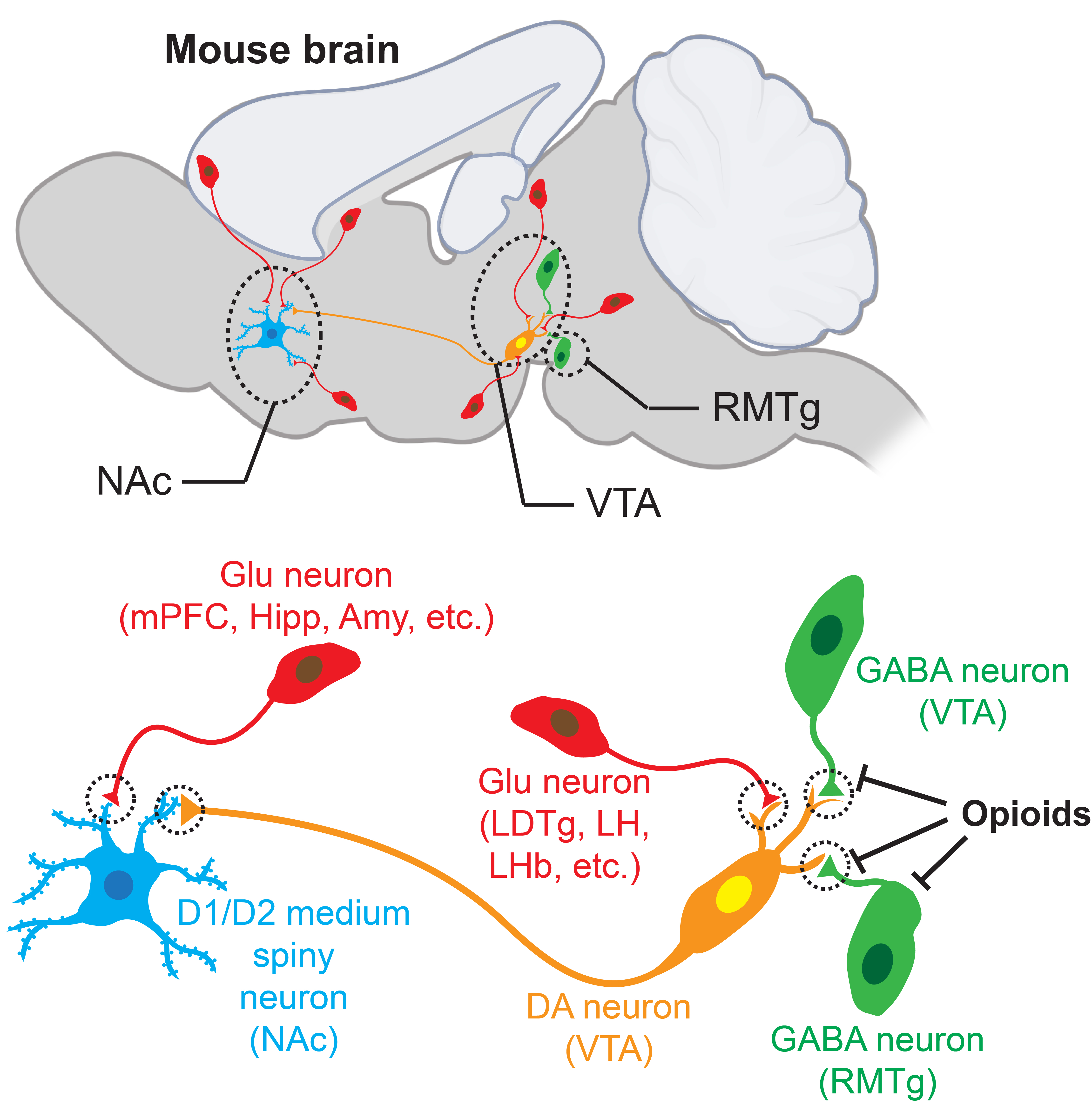
Substance use disorders (SUDs) represent one of the most severe public health crises in the United States. A major barrier to effective treatment is the persistence of relapse vulnerability, which is driven by long-lasting changes in brain circuitry resulting from SUD-induced synaptic plasticity. Building on advances in the field, my lab hypothesizes that synaptic RNA localization and local translation are key mechanisms underlying SUD-induced synaptic plasticity. We aim to develop innovative tools to study this hypothesis using mouse models of SUDs and explore therapeutic strategies to treat SUDs through restoring proper synaptic RNA localization and translation to reverse SUD-induced synaptic plasticity for long-term relapse prevention.
